|
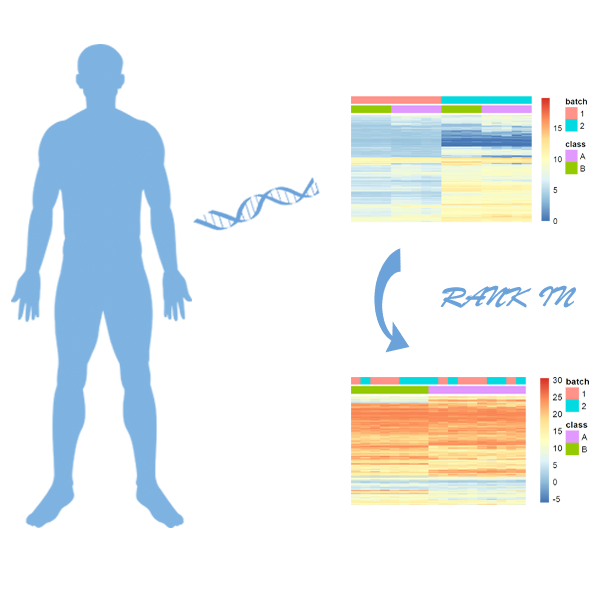
Rank-In is constructed to analyze the consolidated cancer transcriptomics data cross microarray and RNA-seq technologies. Ps: This executable file can be supported by windows 10 system and Python 3.9 |
 |
|
|
Rank-In update:
|
||
|
||
|
Citation: |
||